Prostate cancer is an exceedingly common disease, affecting 1 in 8 men. When diagnosed early, many can live for years without symptoms. Rather than subject early-disease stage patients to the debilitating side-effects of radiation and surgery, clinicians typically monitor these patients through a process called Active Surveillance (AS), using magnetic resonance imaging (MRI) data to track tumor progression, which guides decisions regarding when to begin treatment. Current medical practice uses population level statistics in order to predict and understand how prostate tumors tend to grow. However, this population-based lens may impact detection of patient-specific tumor progression, leading to delayed implementation of life-saving treatment.
New research led by Guillermo Lorenzo, postdoctoral researcher at the Oden Institute of Computational Engineering and Sciences, is catalyzing a revolution in personalized prostate cancer tumor forecasting. Published in Cancer Research Communications, his team’s paper explores how computational models (similarly used by Oden Institute researchers to predict how gliomas and breast cancers will respond to treatment) can be used to predict how tumors develop prior to the treatment stage of this disease. According to Thomas Yankeelov, Oden Institute core faculty and Director of the Center for Computational Oncology, this work “underscores the next generation of tools for guiding the treatment of prostate cancer.”
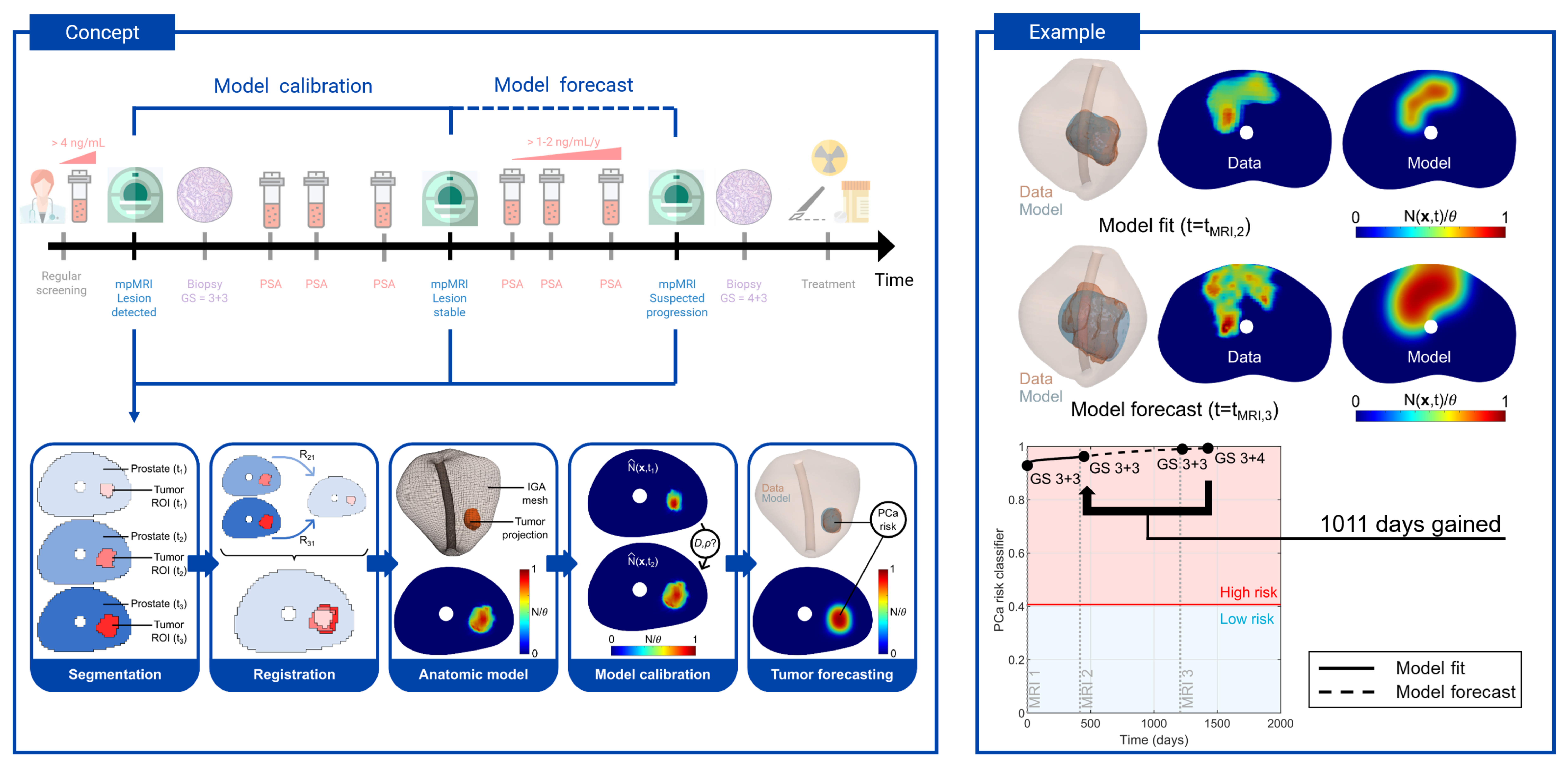
Lorenzo’s team developed a biomechanistic model that uses MRI data from patients’ routine screenings’ during active surveillance to provide personalized tumor forecasting, a coveted capability not yet available in clinical practice. By analyzing cancer cell mobility, which increases tumor size, and the rate of cancer cell division, which increases tumor density, their model is able to computationally and spatiotemporally predict how these interplaying mechanisms will influence future tumor growth within the 3D geometry of a patient’s prostate.
The research team’s technology also allows for the classification and forecasting of dynamic changes in biomarkers that correlate with high-risk tumors. While many prostate cancers begin as low-risk, low-symptom conditions that do not require intervention, they can develop into high-risk cases requiring immediate surgery or radiotherapy. These changes may be missed in the intermittent MRI screenings used in standard-of-care active surveillance. Protocols for detecting increasing cancer risk levels do not account for the vast heterogeneity of prostate cancer growth dynamics across patients.
“Our computational technology for tumor forecasting is a decisive step towards defining a predictive, personalized strategy for the clinical management of prostate cancer that combats the treatment excesses and deficiencies from the current observational, population-based protocols," said Lorenzo.